Protein research
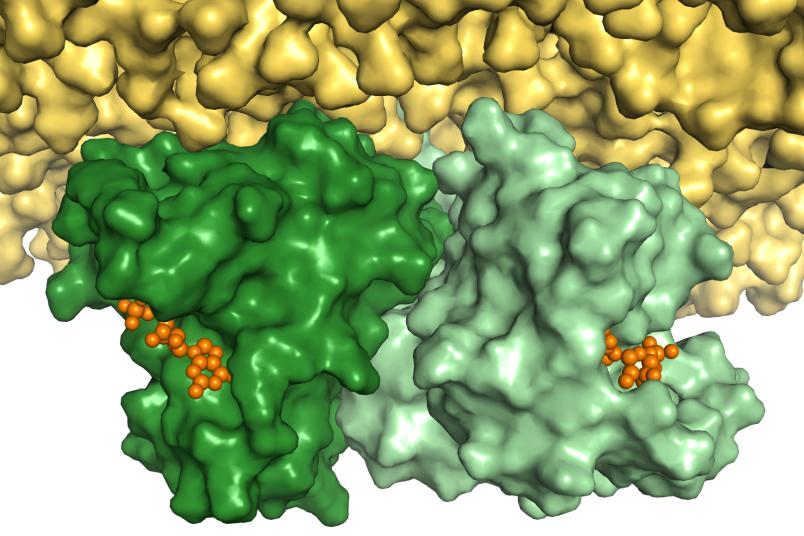
Cancer-promoting Ras protein exists in a pair within cells
Almost all attempts to use the Ras protein as a point of attack for medications have so far failed. New knowledge concerning its structure is raising hopes of options in the development of medications.
Researchers from Bochum and Osnabrück have gained new insights into the structure of the Ras protein, which acts as a molecular switch for cell growth and is involved in the development of cancer. With the help of fluorescence markings, they have demonstrated that the protein is deposited in a pair at the cell membrane, and with the very structure that they predicted in theory back in 2012. The team from the Bochum Center for Protein Diagnostics (PRODI) hopes that these findings will open up a new approach for the development of cancer medications. The researchers from Ruhr-Universität Bochum (RUB) and Osnabrück University published the results in the Journal of Chemical Science in May 2021.
In an activated state, the Ras protein is bound to the molecule guanosine triphosphate, GTP for short. Ras separates one of the phosphate groups from the GTP molecule, thus switching off cell growth. Malfunctions of the Ras protein can cause it to remain in an active state or switch off with a significant delay. This causes uncontrolled cell growth and, as a result, cancer. Almost all attempts to find an active ingredient that docks to individual Ras proteins have so far failed.
Stronger in a pair
Other studies have since shown that the clustering of Ras as a pair causes the signal for cell growth to be transmitted considerably better. “If we could find a way to disrupt the formation of a pair of malfunctioning Ras proteins using an active ingredient, their signal for cell growth would be significantly weakened,” says Professor Klaus Gerwert, Founding Director of PRODI. “As a result, the knowledge of the Ras pair structure offers a new starting point for active ingredients to combat cancer.” Bioinformatic methods could help in the search for potential active ingredients. “However, the precise structure of the interface between the two Ras proteins needs to be known for this,” explains Dr. Till Rudack, lead author of the current study.
Many different contradictory pair models
In 2012, the Bochum group led by Klaus Gerwert identified a structure for the Ras pair at the cell membrane using a specially developed combination of computer-aided and experimental methods. With biomolecular simulations, the group also predicted an initial model of the interface between the two proteins. However, the precise structure of the interface was the subject of controversial debate in the following years and many other contradictory structure models were predicted. “In order to dispel the controversy, experimental data are required to unequivocally validate the model,” says Associate Professor Dr. Carsten Kötting, Project Group Leader at PRODI. These data were provided by the researchers in their current work.
They used the FRET method, short for fluorescence resonance energy transfer, which acts as a kind of molecular yardstick and is used to measure distances between proteins. To do this, two proteins are marked with two different dyes. If they are very close together, energy is transferred from one dye to the other. The distance between the proteins is determined based on the ratio of transferred energy. In order to ascertain the orientation of the two Ras proteins in the pair in relation to each other, the researchers had to measure the distances at different points. “We applied an innovative method to incorporate amino acids into the Ras protein, which are not found naturally in the protein but can attach to the dyes,” explains Carsten Kötting.
Exchange of amino acids destroys the pair
The researchers thus measured a total of four different distances. They then examined with which of the pair models these distances correlated. Their analysis included all of the previously published pair models as well as a further 100 potential models that they predicted by computational methods. However, only one arrangement of the pair matched all four experimentally determined distances. “This arrangement corresponds to the structure we predicted back in 2012,” summarises Till Rudack.
In further experiments, the researchers exchanged two of the most important amino acids at the point of contact between the two Ras proteins. The result: a pair was no longer formed. “The fact that the formation of a pair could be prevented by exchanging amino acids makes me optimistic that we can use the contact point of the Ras pair as a point of attack for active ingredients in the future,” says Klaus Gerwert.